December 8, 2018
Cancer arises when cells lose control. Deciphering the ‘blueprint’ of cancer cells – outlining how cancer cells hijack specific pathways for uncontrolled proliferation – will lead to more efficient ways to fight it.
Joint effort of scientists from the Weizmann Institute of Science and clinicians from major hemato-oncology departments in Israel succeeded in creating detailed profiles of myeloma cancer in both pre-cancer stages, in newly diagnosed multiple myeloma patients as well as post treatment and relapse. These detailed blueprints will help in future precision diagnosis and treatment of this disease.
Multiple myeloma is the second-most common type of blood cancer. Multiple myeloma occurs when plasma cells in the bone marrow – the cells that produce antibodies – proliferate out of control and lead to various type of organ failure and death. Despite many years of research and significant recent improvement in the survival of myeloma patients with new immunotherapy modalities, many patients succumb to limited response and all relapse.
A major stumbling block, in diagnosing myeloma disease is the fact that each patient is unique and current blood tests are incapable of identifying early disease onset and classifying which patient should receive which treatment. For example, patients whose routine blood tests reveal some hallmarks of the disease in an early and precancerous stage are followed closely with a ‘watch and wait’ strategy, but every year 1% of them will lose in this ‘Russian roulette’ and develop the full-blown myeloma disease. Until recently, there had been no way to discriminate between those that will transform and those that will not.
Dr Guy Ledergor, an MD/PhD and Dr Assaf Weiner, in Professor Ido Amit’s group from the Weizmann Institute of Science’s Immunology Department, together with Professor Amos Tanay of the Biological Regulation and Computer Science and Applied Mathematics Departments, thought that a highly sensitive method developed by the group, known as single cell RNA sequencing, may provide a new approach to understanding the makeup of multiple myeloma and devise new and more effective approaches to diagnose and treat this devastating disease. With this vision in mind, The Weizmann Institute of Science’s Bench to Bedside Program, headed by Professor Gabi Barbash, facilitated an Israeli nation-wide effort towards recruiting all of the hemato-oncology departments in Israel.
The new method sequences the RNA in thousands of individual cells from the patient blood or bone marrow, allowing to capture the specific gene program that is active in each individual cell.
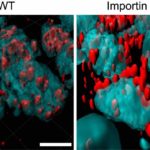
In order to understand the myeloma cancer blueprint, the scientists first generated a high-resolution model of normal plasma cells by sequencing tens of thousands of cells from healthy individuals undergoing hip-replacements who served as a control group. The plasma cells of the control patients were highly similar within and across individuals, basically showing a single, common blue print of normal plasma cells.
Comparing the normal blueprint to the blueprints of myeloma patients, showed that the blueprint of myeloma cancer cells are extremely heterogeneous, with every patient showing its own, unique blueprint and with some patients demonstrating several tumour clones displaying more than a single blueprint in one patient.
This joint effort by clinicians and scientists from the Weizmann Institute of Science has been able to demonstrate that it is possible to identify even a very small number of malignant cells in the blood at very early (pre-cancer) stage, and allowing much more precise diagnosis and informed choices about treatments for each patient based on his ‘personalised’ disease.
As a result, it will also be possible to follow and monitor these patients with significantly less painful blood tests, replacing today’s painful bone marrow biopsies. Eventually the method might help multiple myeloma patients receive earlier and more precise treatments, and possibly prevent the relapse that often follows chemotherapy.
This study and the sensitive method used for the first time world-wide, is now published in Nature Medicine and may help to advance Israel to the forefront of myeloma patient diagnosis and treatment.
Professor Barbash points out that these findings are just the beginning for applying single cell RNA-seq technologies to clinical-genomic research.
“We are now developing similar collaborations between hospital-based researchers and basic-research scientists to advance diagnosis and treatment for other maladies,” he said.
“Single cell genomic analysis until now was confined to a small number of research labs, we are constantly pushing the boundaries of the technology in ways that will make it a major clinical discovery and diagnostics tool,” said Professor Amit.
Such tracking may be useful for diagnosing early signs of many malignancies in the precancerous stage or post-chemotherapy relapse, thus becoming part of a personalised approach. It will enable decisions to be made based on the individual profiles of each patient’s cells at every stage of disease. These profiles are already used to create targeted therapies against the cancer cells.
The researchers used a machine-learning approaches to automatically identify malignant cells out of tens of thousands of cells.
“We are entering an era in which measuring big-data and using machine learning will provide clinicians new insights and understanding into devastating diseases like multiple myeloma,” said Dr Weiner.
“In the future, physicians will hopefully be able to track the disease in real time and treat each patient according to a personalised disease profile, possibly even before symptoms appear,” said Professor Ledergor.
The story has been so far covered in the following media outlets:
BREAKINGISRAELNEWS SCIENCE DAILY MEDKIT REACHMD MEDICALXPRESS BIOENGINEER
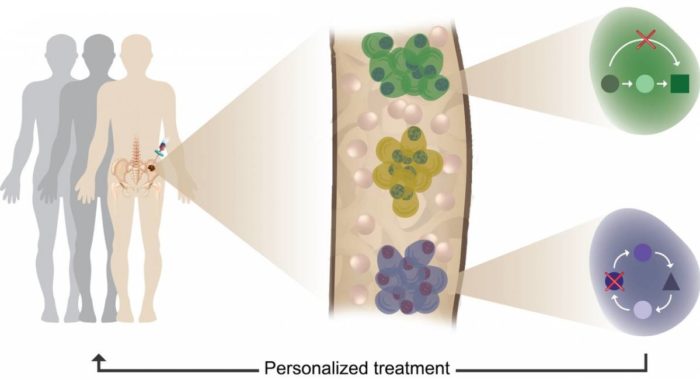
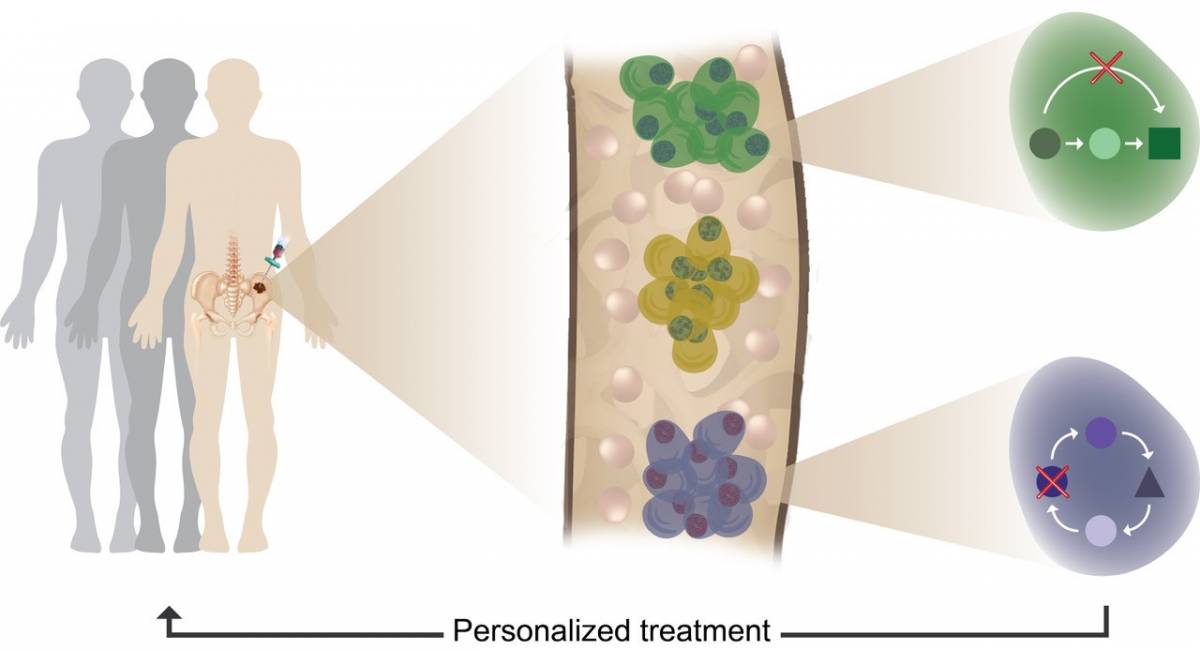
Tumour heterogeneity, clonal evolution and therapy resistance revealed using single cell profiling of multiple myeloma patients. Illustration: Noa Godin