November 17, 2018
A computational method for designing dramatically more efficient versions of enzymes, being developed at the Weizmann Institute of Research, may lead to new avenues of drug design and new treatments against nerve agents.
Millions of times a second, in our bodies and in every living organism, molecules are being cut up or stuck together, thanks to enzymes.
Enzymes are proteins equipped for various tasks, but natural ones may not always be as efficient as we would like. For example, the enzymes that break down toxins must be stable long enough to carry out their jobs and they need to be quick, efficient and precise.
Variations in these properties of an enzyme can mean life or death for the organism, and while some variations may be lethal, others might grant an advantage to the plant or animal. But could we identify the variations – mutations in the genetic sequences encoding the proteins’ structure – and use them to create better enzymes? The problem is that a true improvement is likely to involve a combination of several mutations, and each alone might only increase the efficiency minimally. The number of possible combinations of mutations for any given enzyme is astronomical, and there would be no way to test them all.
Dr Sarel Fleishman of the Biomolecular Sciences Department of the Weizmann Institute of Science and his research group are developing a way around this problem. The automatic computerised system they created can look for the small changes that might lead to significant increases in activity and identify those versions that are likely to work at very high levels of efficiency. This system, they hope, may lead to significant developments in biotechnology and medicine, as it essentially takes the guesswork out of redesigning proteins for particular tasks.
Fleishman’s group, together with that of Professor Dan Tawfik, also of the Biomolecular Sciences Department, recently demonstrated the power of this method by focusing on a particular enzyme – one that breaks down molecules of nerve agents such as sarin and VX.
The natural version of this enzyme is sadly inefficient against these neurotoxins, but Tawfik and Fleishman thought that the enzyme could be remodelled to become more efficient so that it could be used either as an antidote or as a clean-up for affected sites. Tawfik and his group have been using directed evolution – the method for which was awarded the 2018 Nobel Prize in Chemistry – to reprogram the natural enzyme so that it would detoxify nerve agents. But directed evolution is a laborious process involving the screening thousands of enzyme mutants to identify a few improved ones.
The method developed by Fleishman and his team offered a new way to approach the issue: Staff Scientist Dr Olga Khersonsky, and Rosalie Lipsh and Ziv Avizemer, research students in his group, took up the challenge.
The automated system created dozens of new versions of this enzyme, each bearing up to five mutations on top of the natural gene. This produced several designer versions of the enzyme that proved to be up to 4,000 times better than the original at breaking down nerve agents.
“That is an enormous improvement,” said Fleishman.
“If you could somehow reproduce that level of improvement in an average human runner, he would make it from Rehovot to Haifa (about 125 km) before I finished uttering this sentence.”
The new computerised protein revision method is applicable to nearly all of the many proteins that are candidates for optimization. Fleishman and his team have also developed a website that any researcher can use. They only need simply supply the information they have on the protein in question, and several hours later they can obtain a limited number of possible candidates for testing.
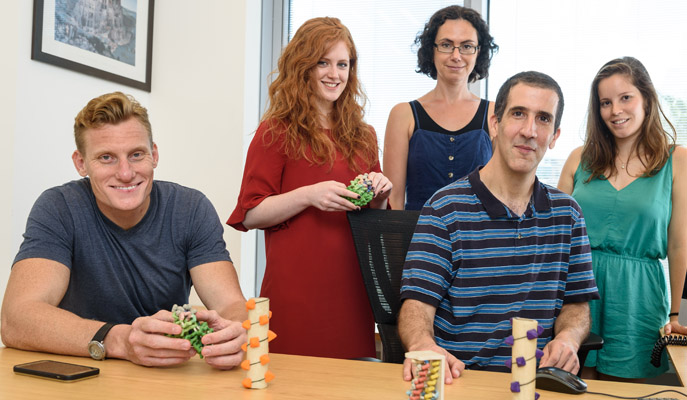
The designed version of this protein contains numerous tiny mutations that, added together, make it thousands of time better at breaking down nerve agents like sarin or VX

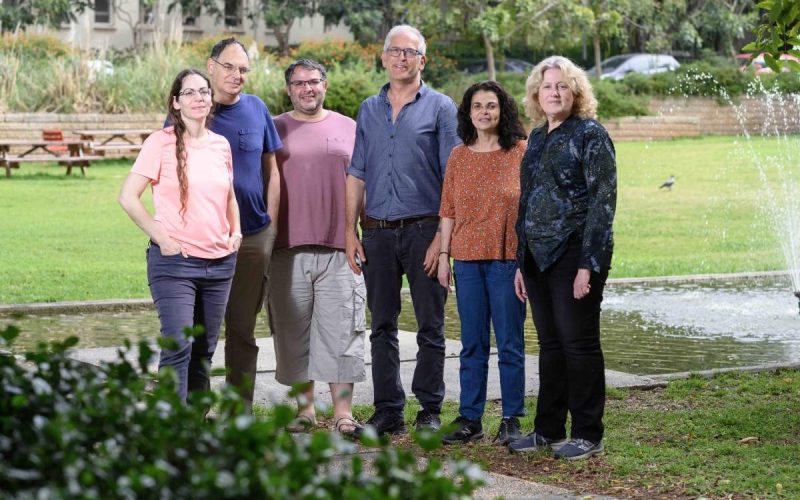
(l-r) Protein planners Dr Gideon Lapidoth, Rosalie Lipsh, Dr Olga Khersonsky, Dr Sarel Fleishman and Ziv Avizemer

Professor Dan Tawfik





